New preprint: Metagenomic sequencing of composite airplane wastewater for surveillance of emerging viruses
Note: The Nucleic Acid Observatory (NAO) is now SecureBio Detection.
This post describes work done with Olivia Hershey, Ari Machtinger, Dan Rice, Will Bradshaw, and collaborators from Ginkgo Biosecurity and the CDC. Thanks to Lenni Justen, Dan Rice, and Jeff Kaufman for feedback on the post.
Our interest in airplane wastewater for pathogen detection began with its ability to target long-distance travelers, who can quickly spread new pathogens around the globe. However, we knew from past work that metagenomics-based detection can depend less on who you sample than on how much the viral signal is obscured by other microbes—specifically the bacteria that thrive in municipal sewer systems. In a study with collaborators from Ginkgo Biosecurity and the CDC, we found that human viruses typically had around 13-fold higher relative abundance in composite (pooled) airplane wastewater than municipal wastewater collected from a nearby treatment plant. This systematic increase appears to be largely driven by less sewer-derived bacteria being present in airplane samples. These findings suggest that composite airplane wastewater can be a particularly cost-effective sample type for detecting emerging viruses. We’ve shared these findings as a preprint, “Metagenomic sequencing of composite airplane wastewater for surveillance of emerging viruses”, and give a more detailed summary below.
Why monitor air travelers?
Long-distance air travelers play a key role in the global spread of infectious pathogens. During the early stages of an outbreak, infected travelers can carry a virus from its origin to cities around the world within days. This makes the traveler population a valuable sentinel for pandemic early warning: Detecting a novel pathogen circulating among travelers could provide advance notice before it becomes established in local communities.
Composite airplane wastewater can provide a non-invasive, cost-effective way to monitor disease among this population. Sampling individual flights is expensive and logistically challenging; however, pooled airplane waste can be sampled relatively easily at airplane waste triturators, devices used by airports to process airplane waste before it enters the sewer system. An autosampling device connected to a triturator can siphon small amounts of airplane wastewater into a single composite sample, mirroring the approach commonly used for sampling wastewater at municipal treatment plants. Airports typically split all airplane waste across a small number of triturators, such that a 24-hour composite sample collected at a triturator will contain wastewater from a large number of the past day’s incoming flights.
The challenge of metagenomic sequencing in wastewater
Current wastewater surveillance programs primarily use targeted assays like qPCR, which can only detect pathogens we already know to look for. Metagenomic sequencing takes a different approach: By sequencing all the genetic material in a sample, it can in principle detect any virus, including ones that have never been seen before. But human viruses make up only a tiny fraction of the RNA in wastewater (roughly 1 in 10,000 sequences generated by CASPER). The vast majority comes from bacteria, much of it from microbes that live not in humans but in the sewer system itself.1 The low relative abundance2 of human viruses makes detection expensive, as you need to generate enormous amounts of data to reliably see them (Grimm et al. (2025)).
What we know about wastewater metagenomics primarily comes from studies of wastewater from municipal treatment plants. Airplane wastewater takes a much shorter path from excretion to sampling compared to the miles of biofilm-laden sewer pipes that municipal waste must pass through. Airplane waste holds are also subject to cold temperatures and filled with disinfectant fluid which may inhibit microbial activity. All of these differences plausibly reduce the influx and growth of sewer-derived microbes and/or help preserve human-derived viruses, thereby increasing the relative abundance of the viruses we care about.
An experiment to evaluate metagenomic sequencing of composite airplane wastewater
To evaluate the utility of composite airplane wastewater for viral metagenomics, we worked with Ginkgo Biosecurity and the CDC to collect samples of composite airplane wastewater over two months in late 2023 from a triturator at Boston Logan International Airport (as part of the CDC’s Traveler-based Genomic Surveillance program). In parallel, we collected samples of municipal wastewater from Deer Island Treatment Plant, which serves the Greater Boston area. We processed samples using a protocol designed to enrich for viruses over bacteria, then performed untargeted viral metagenomic sequencing. Here, we report on an analysis of sequencing data from a subset of 4 days across an 11-day period in mid-December 2023, focusing on how the relative abundance of various human viruses differs between wastewater from the triturator versus the treatment plant.
Human viruses have higher relative abundance in airplane wastewater
Across the ten genera of human-infecting RNA viruses that we examined, relative abundances were consistently higher in airplane wastewater than in municipal wastewater. The median increase was 13-fold, with a range from about 5.5-fold for Sapovirus to 120-fold for Alphacoronavirus (Figure 1, top section). These groups span diverse viral families and include both respiratory viruses (influenza, coronaviruses) and enteric viruses (norovirus, rotavirus), suggesting that the increase is systematic rather than specific to particular pathogens.
We saw a similar pattern for other human-associated organisms. Dietary plant viruses like pepper mild mottle virus, human gut bacteria like Bacteroides, and even human sequences all showed elevated relative abundances in airplane samples (Figure 1, middle section). Conversely, bacteria known to be associated with sewer infrastructure, such as Arcobacter and Trichococcus, were typically reduced (Figure 1, bottom section).
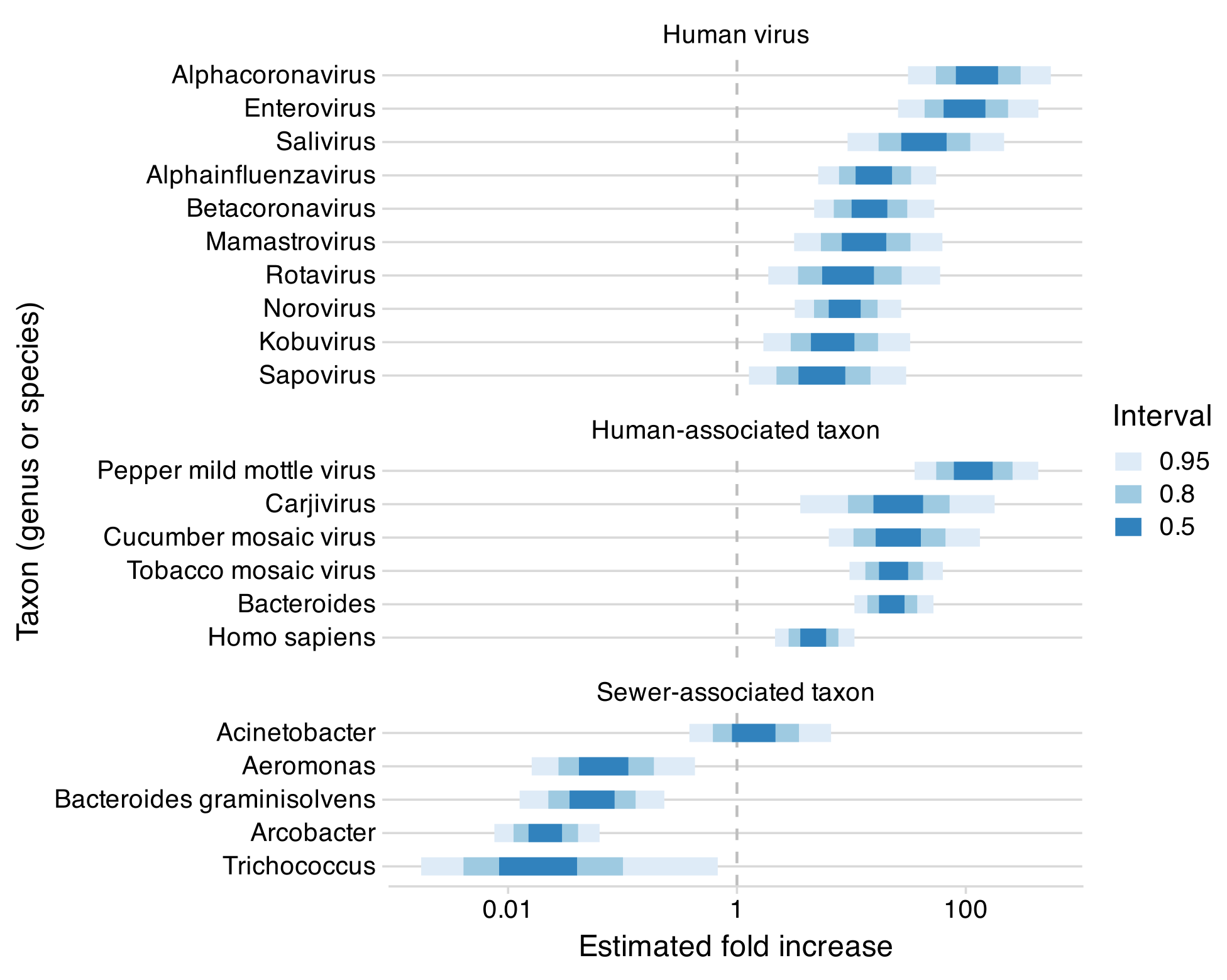
Figure 1: The estimated fold increase in the relative abundance of various biological taxa (groups of organisms such as a genus or species) in airplane versus municipal wastewater (Figure 4 from the preprint). Human viruses (top) and other human-associated taxa (middle) show increases much greater than one, while most sewer-associated bacteria (bottom) show decreases. Bars show 50%, 80%, and 95% credible intervals from a Bayesian negative binomial regression model.
This overall pattern can be explained by processed samples of airplane wastewater having a higher ratio of human- to sewer-derived genetic material compared to samples of municipal wastewater. This explanation implies that the large increase in relative abundances seen for known human RNA viruses would also apply to a novel one.
What this means for biosurveillance costs
Higher viral relative abundances translate directly to lower sequencing costs. All else equal, if a virus is 10 times more abundant, you need roughly one-tenth the sequencing to detect it and/or characterize its genome.3 For example, if 1 in 1,000 people are infected with SARS-CoV-2, we estimate that detecting the virus via metagenomics in a sample of municipal wastewater would require roughly $1,000 worth of sequencing.4 With the 13-fold increase we observed in airplane wastewater, that drops to around $75. While targeted assays like qPCR remain cheaper for any one virus, this reduction in cost applies simultaneously to all viruses covered by the metagenomics assay, including undiscovered ones for which no targeted assay exists.
Limitations and future directions
This study analyzes sequencing data from just an 11-day period. We have sequenced a longer period (roughly 6 weeks) which can provide insight into how these differences vary over time and whether airplane wastewater typically presages local outbreaks. This study was limited to a single airport and treatment plant over one season, so multi-site studies will be needed to assess how these findings generalize.
Higher average relative abundances mean we are much more likely to detect a virus when an infected person contributes to an airplane sample. However, even composite airplane samples covering many flights are likely to contain material from far fewer people than a sample from a large treatment plant. As a result, airplane samples may be more likely to miss rare infections. More work is needed to assess how this tradeoff impacts the overall sensitivity of metagenomics-based detection.
Conclusion
Collecting composite airplane wastewater at airport triturators provides a streamlined way to monitor the traveler populations that drive global pathogen spread. The higher viral relative abundances we observed in airplane (versus municipal) wastewater may substantially reduce the cost of effective metagenomic detection, providing a scalable path to a pathogen-agnostic early warning system.
Read the full preprint at “Metagenomic sequencing of composite airplane wastewater for surveillance of emerging viruses”.
References
Fierer, Noah, Hannah Holland-Moritz, Alexandra Alexiev, Harpreet Batther, Nicholas B. Dragone, Liam Friar, Matthew J. Gebert, et al. 2022. “A Metagenomic Investigation of Spatial and Temporal Changes in Sewage Microbiomes Across a University Campus.” mSystems 7 (5): e00651–22. https://doi.org/10.1128/msystems.00651-22.
Grimm, Simon L., Jeff T. Kaufman, Daniel P. Rice, Charles Whittaker, William J. Bradshaw, and Michael R. McLaren. 2025. “Inferring the Sensitivity of Wastewater Metagenomic Sequencing for Early Detection of Viruses: A Statistical Modelling Study.” The Lancet Microbe 6 (11): 101187. https://doi.org/10.1016/j.lanmic.2025.101187.
McLellan, Sandra L, and Adélaïde Roguet. 2019. “The Unexpected Habitat in Sewer Pipes for the Propagation of Microbial Communities and Their Imprint on Urban Waters.” Current Opinion in Biotechnology, Energy Biotechnology • Environmental Biotechnology, 57 (June): 34–41. https://doi.org/10.1016/j.copbio.2018.12.010.
Newton, Ryan J., Sandra L. McLellan, Deborah K. Dila, Joseph H. Vineis, Hilary G. Morrison, A. Murat Eren, and Mitchell L. Sogin. 2015. “Sewage Reflects the Microbiomes of Human Populations.” mBio 6 (2): 10.1128/mbio.02574–14. https://doi.org/10.1128/mbio.02574-14.
Roguet, Adélaïde, Ryan J. Newton, A. Murat Eren, and Sandra L. McLellan. 2022. “Guts of the Urban Ecosystem: Microbial Ecology of Sewer Infrastructure.” mSystems 7 (4): e00118–22. https://doi.org/10.1128/msystems.00118-22.
VandeWalle, J. L., G. W. Goetz, S. M. Huse, H. G. Morrison, M. L. Sogin, R. G. Hoffmann, K. Yan, and S. L. McLellan. 2012. “Acinetobacter, Aeromonas and Trichococcus Populations Dominate the Microbial Community Within Urban Sewer Infrastructure.” Environmental Microbiology 14 (9): 2538–52. https://doi.org/10.1111/j.1462-2920.2012.02757.x.